|
Genecopoeia
hek293t cells Hek293t Cells, supplied by Genecopoeia, used in various techniques. Bioz Stars score: 96/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/hek293t cells/product/Genecopoeia Average 96 stars, based on 1 article reviews
hek293t cells - by Bioz Stars,
2026-03
96/100 stars
|
Buy from Supplier |
|
R&D Systems
goat anti ace2 antibody Goat Anti Ace2 Antibody, supplied by R&D Systems, used in various techniques. Bioz Stars score: 99/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/goat anti ace2 antibody/product/R&D Systems Average 99 stars, based on 1 article reviews
goat anti ace2 antibody - by Bioz Stars,
2026-03
99/100 stars
|
Buy from Supplier |
|
R&D Systems
fluorescence conjugated antibodies targeting ace2 Fluorescence Conjugated Antibodies Targeting Ace2, supplied by R&D Systems, used in various techniques. Bioz Stars score: 91/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/fluorescence conjugated antibodies targeting ace2/product/R&D Systems Average 91 stars, based on 1 article reviews
fluorescence conjugated antibodies targeting ace2 - by Bioz Stars,
2026-03
91/100 stars
|
Buy from Supplier |
|
Sino Biological
human ace2 protein  Human Ace2 Protein, supplied by Sino Biological, used in various techniques. Bioz Stars score: 96/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/human ace2 protein/product/Sino Biological Average 96 stars, based on 1 article reviews
human ace2 protein - by Bioz Stars,
2026-03
96/100 stars
|
Buy from Supplier |
|
Addgene inc
prrl sin cppt sffv ace2 ires puro wpre  Prrl Sin Cppt Sffv Ace2 Ires Puro Wpre, supplied by Addgene inc, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/prrl sin cppt sffv ace2 ires puro wpre/product/Addgene inc Average 93 stars, based on 1 article reviews
prrl sin cppt sffv ace2 ires puro wpre - by Bioz Stars,
2026-03
93/100 stars
|
Buy from Supplier |
|
Sino Biological
human ace2  Human Ace2, supplied by Sino Biological, used in various techniques. Bioz Stars score: 96/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/human ace2/product/Sino Biological Average 96 stars, based on 1 article reviews
human ace2 - by Bioz Stars,
2026-03
96/100 stars
|
Buy from Supplier |
|
BPS Bioscience
ace2 hek293 recombinant cell line  Ace2 Hek293 Recombinant Cell Line, supplied by BPS Bioscience, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/ace2 hek293 recombinant cell line/product/BPS Bioscience Average 93 stars, based on 1 article reviews
ace2 hek293 recombinant cell line - by Bioz Stars,
2026-03
93/100 stars
|
Buy from Supplier |
|
Proteintech
66 699 1 ig  66 699 1 Ig, supplied by Proteintech, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/66 699 1 ig/product/Proteintech Average 94 stars, based on 1 article reviews
66 699 1 ig - by Bioz Stars,
2026-03
94/100 stars
|
Buy from Supplier |
|
Genecopoeia
human ace2  Human Ace2, supplied by Genecopoeia, used in various techniques. Bioz Stars score: 94/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/human ace2/product/Genecopoeia Average 94 stars, based on 1 article reviews
human ace2 - by Bioz Stars,
2026-03
94/100 stars
|
Buy from Supplier |
|
Addgene inc
tmprss2 expression plasmids  Tmprss2 Expression Plasmids, supplied by Addgene inc, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/tmprss2 expression plasmids/product/Addgene inc Average 93 stars, based on 1 article reviews
tmprss2 expression plasmids - by Bioz Stars,
2026-03
93/100 stars
|
Buy from Supplier |
|
Addgene inc
rrl sin cppt sffv ace2 ires neo wpre mt129 addgene 145840  Rrl Sin Cppt Sffv Ace2 Ires Neo Wpre Mt129 Addgene 145840, supplied by Addgene inc, used in various techniques. Bioz Stars score: 93/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/rrl sin cppt sffv ace2 ires neo wpre mt129 addgene 145840/product/Addgene inc Average 93 stars, based on 1 article reviews
rrl sin cppt sffv ace2 ires neo wpre mt129 addgene 145840 - by Bioz Stars,
2026-03
93/100 stars
|
Buy from Supplier |
|
Thermo Fisher
gene exp ace2 hs01085340 m1  Gene Exp Ace2 Hs01085340 M1, supplied by Thermo Fisher, used in various techniques. Bioz Stars score: 90/100, based on 1 PubMed citations. ZERO BIAS - scores, article reviews, protocol conditions and more https://www.bioz.com/result/gene exp ace2 hs01085340 m1/product/Thermo Fisher Average 90 stars, based on 1 article reviews
gene exp ace2 hs01085340 m1 - by Bioz Stars,
2026-03
90/100 stars
|
Buy from Supplier |
Image Search Results

Journal: Journal of Chemical Theory and Computation
Article Title: Differential Interactions between Human ACE2 and Spike RBD of SARS-CoV-2 Variants of Concern
doi: 10.1021/acs.jctc.1c00965
Figure Lengend Snippet: (A) Average force profiles of WT (red), Alpha (blue), Beta (orange), Gamma (sky blue), Epsilon (green), Kappa (pink), and Delta (gray) variants as a function of the distance between the centers of mass of RBD and ACE2. (B) Initial snapshot of WT. Residues subjected to each mutation are shown as solid sticks (N501, K417, E484, L452, and T478). RBD and ACE2 are, respectively, colored in light gray and yellow. All N-glycans, water, and ions are hidden for clarity. (C) Initial snapshot of WT with clockwise 90° rotation along the normal from (B). All N-glycans are depicted in different colors. Any other residues, water, and ions are not shown for clarity.
Article Snippet: The recombinant
Techniques: Mutagenesis

Journal: Journal of Chemical Theory and Computation
Article Title: Differential Interactions between Human ACE2 and Spike RBD of SARS-CoV-2 Variants of Concern
doi: 10.1021/acs.jctc.1c00965
Figure Lengend Snippet: Two-dimensional contact maps at D = 53 Å. (A) Interacting residue pairs between RBD WT and ACE2. RBD residues subjected to mutation are shown in colored boxes at the bottom: (B) blue for Alpha, (C) orange for Beta, and (D) green for Epsilon. The contact frequency is numbered with colors from light blue to dark blue. Dark red and yellow colors on the map, respectively, represent increased and decreased interactions between RBD and ACE2 upon mutations.
Article Snippet: The recombinant
Techniques: Mutagenesis

Journal: Journal of Chemical Theory and Computation
Article Title: Differential Interactions between Human ACE2 and Spike RBD of SARS-CoV-2 Variants of Concern
doi: 10.1021/acs.jctc.1c00965
Figure Lengend Snippet: (A) The average number of contacts between RBD residue 501 and ACE2. (B, C) Representative snapshots at D = 53 Å of (B) Alpha variant and (C) WT. (D) Average number of contacts between RBD residue 417 and ACE2 and (E, F) their interacting residue pairs at D = 53 Å of (E) Beta and (F) Alpha variants. (G) Average number of contacts between RBD residue 478 and ACE2 and (H, I) key interaction pairs at D = 78 Å of (H) Delta and (I) Epsilon variants. The overall color scheme is the same as in Figure , and each mutated residue in each variant is shown using the same colors (i.e., red for WT, blue for Alpha, orange for Beta, green for Epsilon, and gray for Delta). Interacting residues are depicted as solid sticks, and residues losing their interactions are shown as transparent sticks. RBD and ACE2 are presented in light gray and yellow, respectively.
Article Snippet: The recombinant
Techniques: Variant Assay

Journal: Journal of Chemical Theory and Computation
Article Title: Differential Interactions between Human ACE2 and Spike RBD of SARS-CoV-2 Variants of Concern
doi: 10.1021/acs.jctc.1c00965
Figure Lengend Snippet: Binding affinities between RBD variants and ACE2 and its comparison with the simulation results. K d is obtained from microscale thermophoresis experiments. F WT / F is a ratio, where F WT and F are the respective maximum pulling force of WT and of each variant obtained from the SMD simulations.
Article Snippet: The recombinant
Techniques: Binding Assay, Microscale Thermophoresis, Variant Assay

Journal: Immunology and Cell Biology
Article Title: The vaccinia‐based Sementis Copenhagen Vector coronavirus disease 2019 vaccine induces broad and durable cellular and humoral immune responses
doi: 10.1111/imcb.12539
Figure Lengend Snippet: Spike‐specific antibody and SARS‐CoV‐2 neutralization responses following SCV‐S vaccination. (a) S1 and S2 subunit endpoint IgM and IgG ELISA titers determined from serum of female C57BL/6J mice ( n = 3) at the indicated times after a single vaccination with 10 7 PFU of SCV‐S, with (b) ratio of S1‐ and S2‐specific IgG2c to IgG1 endpoint ELISA titers determined 28 days after vaccination. (c) S1 IgG ELISA binding titers (left panel) and cPass neutralization titers (right panel) in outbred ARC(s) and inbred C57BL/6J female mice ( n = 5) 21 days after a single vaccination with 10 7 PFU of SCV‐S or vector control, with (d) durability of response in the outbred cohort shown by S1‐specific endpoint IgG ELISA titers (left panel) and neutralization titers (right panel) at the indicated times. (e) S1‐specific IgG ELISA (left panel) and neutralization titers (right panel) 50 days after a single‐dose (day 0) or prime‐boost (day 0 and 28) vaccination of female C57BL/6J mice ( n = 5) with 10 7 PFU SCV‐S or control vector, with (f) neutralizing activity in ACE2 and RBD blocking ELISA, and (g) neutralizing activity (IC 80 titers) against lenti‐SARS‐CoV‐2‐S pseudoviruses bearing spike protein from the Wuhan reference strain, the alpha or beta variant, shown for prime‐boost samples. (h) Neutralization titers in young (6–8 weeks old; n = 5) and middle‐aged (9–10 months old; n = 10) C57BL/6J mice at the indicated times after prime‐boost vaccination with 10 7 PFU SCV‐S. Results shown are representative of four independent experiments (indicated above) with binding and neutralizing antibody levels comparable at similar doses and time points across all experiments. Symbols represent individual mice and bars show the mean ± s.e.m. from independent experiments. Data were log transformed and statistical significance determined using Brown–Forsythe and Welch ANOVA with Dunnett T3 multiple comparison test. * P < 0.05; ** P < 0.01; *** P < 0.001; **** P < 0.0001. ACE2, angiotensin‐converting enzyme‐2; IC 80 , 80% inhibitory concentration; Ig, immunoglobulin; ns, not significant; PFU, plaque‐forming units; RBD, receptor‐binding domain; SARS‐CoV‐2, severe acute respiratory syndrome coronavirus 2; SCV‐S, Sementis Copenhagen Vector spike protein.
Article Snippet:
Techniques: Neutralization, Enzyme-linked Immunosorbent Assay, Binding Assay, Plasmid Preparation, Activity Assay, Blocking Assay, Variant Assay, Transformation Assay, Concentration Assay

Journal: bioRxiv
Article Title: SARS-CoV-2 Nsp15 antagonizes the cGAS-STING-mediated antiviral innate immune responses
doi: 10.1101/2024.09.05.611469
Figure Lengend Snippet: (A) Schematic diagram of the recombinant SARS-CoV-2 genome with Nsp15 H234A and N277A mutations. (B) Replication kinetics of Nsp15 WT , Nsp15 H234A , and Nsp15 N277A rSARS-CoV-2 in Vero E6 cells (MOI of 0.001). (C) Cell lysates from mock- and virus-infected Vero E6 cells (MOI of 0.001) after 48 h of infection were incubated with anti-Nsp15 antibody or isotype control. Immunoblot analysis applied for the immunoprecipitated Nsp15 proteins. (D) Western blot for ACE2 and STAT1 in A549-ACE2 cells with or without STAT1 knockout. (E and F) Replication kinetics of Nsp15 WT , Nsp15 H234A , and Nsp15 N277A rSARS-CoV-2 in A549-ACE2 and A549-ACE2/STAT1 KO cells (MOI of 1). (G) The area under the curve (AUC) was measured from each viral growth curve and plotted as a bar graph. Data are mean ± SD (n = 3) and analyzed by two-way ANOVA with Dunnett’s multiple comparison test. (H-K) A549-ACE2 cells were uninfected or infected by Nsp15 WT , Nsp15 H234A , and Nsp15 N277A rSARS-CoV-2 at MOI of 1 for 8 and 24 h. Total RNA collected for evaluating the host responses by RT-qPCR. Relative gene expression was normalized by 18S rRNA and presented relative to mock infection. Data are mean ± SD (n = 3) and analyzed by two-way ANOVA with Dunnett’s multiple comparison test. * P ≤0.05; *** P ≤0.001; **** P ≤0.0001; ns, not significant.
Article Snippet: BHK-21-hACE2 cell, a derivative of BHK-21 cell (ATCC, CCL-10), was established by transduction of lentiviral particle bearing
Techniques: Recombinant, Virus, Infection, Incubation, Control, Western Blot, Immunoprecipitation, Knock-Out, Comparison, Quantitative RT-PCR, Gene Expression

Journal: bioRxiv
Article Title: SARS-CoV-2 Nsp15 antagonizes the cGAS-STING-mediated antiviral innate immune responses
doi: 10.1101/2024.09.05.611469
Figure Lengend Snippet: (A) Design of rVSV-EGFP genome bearing SARS-CoV-2 Nsp15. (B) Vero cells were uninfected or infected by rVSV-EGFP expressing WT and H234A Nsp15 at MOI of 0.1 for 16 h. Western blot performed to verify the expression of indicated proteins. (C and D) Replication kinetics of parental rVSV-EGFP and rVSV-EGFP expressing WT and H234A Nsp15 in A549-ACE2 and A549-ACE2/STAT1 KO cells (MOI of 0.1). Data are mean ± SD (n = 3) and analyzed by two-way ANOVA with Dunnett’s multiple comparison test. (E-H) Total RNA collected from A549-ACE2 cells with rVSV-Nsp15 WT and -Nsp15 H234A infection and mock infection. RNA level of indicated host genes relative to 18S rRNA was measured by RT-qPCR and presented by fold change over mock infection. Data are mean ± SD (n = 3) and analyzed by two-way ANOVA. * P ≤0.05; ** P ≤0.01; *** P ≤0.001.
Article Snippet: BHK-21-hACE2 cell, a derivative of BHK-21 cell (ATCC, CCL-10), was established by transduction of lentiviral particle bearing
Techniques: Infection, Expressing, Western Blot, Comparison, Quantitative RT-PCR

Journal: bioRxiv
Article Title: SARS-CoV-2 Nsp15 antagonizes the cGAS-STING-mediated antiviral innate immune responses
doi: 10.1101/2024.09.05.611469
Figure Lengend Snippet: A549-ACE2 cells with mock, Nsp15 WT , Nsp15 H234A , and Nsp15 N277A rSARS-CoV-2 infection for 8 and 24 h (MOI of 5). Total RNA with poly(A) enrichment followed by RNA sequencing analysis. (A) Schematic of bulk RNA-seq experimental design (n = 3 per group). (B) PCA of total normalized transcript abundance from mock, Nsp15 wild-type and mutant rSARS-CoV-2 infection. Sparse PCA depicts the global transcriptome of individual sample. (C) Venn diagram for unique and shared differentially expressed genes (Padj < 0.05 & |log2FC| > 1) in cells infected with Nsp15 H234A and Nsp15 N277A mutants compared to Nsp15 WT virus. (D) Volcano plots showing GSEA results generated using MSigDB Hallmark pathways. (E) Cluster heatmap of GSVA scores generated using representative innate immune and metabolic signatures from MSigDB Gene Ontology signatures. (F) Expression heatmap of representative innate immune and metabolic genes in across mock, wild-type, and mutant rSARS-CoV-2 infection.
Article Snippet: BHK-21-hACE2 cell, a derivative of BHK-21 cell (ATCC, CCL-10), was established by transduction of lentiviral particle bearing
Techniques: Infection, RNA Sequencing, Mutagenesis, Virus, Generated, Expressing

Journal: bioRxiv
Article Title: SARS-CoV-2 Nsp15 antagonizes the cGAS-STING-mediated antiviral innate immune responses
doi: 10.1101/2024.09.05.611469
Figure Lengend Snippet: (A and B) Endogenous cGAS and STING mRNA levels in A549-ACE2 cells uninfected or infected with Nsp15 WT and Nsp15 H234A rSARS-CoV-2 at MOI of 5 for 8 or 24 h. RT-qPCR results are presented relative to the expression of 18S rRNA. Data are mean ± SD (n = 3) and analyzed by two-way ANOVA with Tukey’s multiple comparison test. * P ≤0.05; **** P ≤0.0001; ns, not significant. (C) BHK-ACE2 cells were mock infected or infected with Nsp15 WT and Nsp15 H234A rSARS-CoV-2 at MOI of 0.1, further transfected with cGAS and STING plasmids at 4 hpi. At 24 and 48 hpi, cell lysates were harvested for clarifying the indicated protein levels.
Article Snippet: BHK-21-hACE2 cell, a derivative of BHK-21 cell (ATCC, CCL-10), was established by transduction of lentiviral particle bearing
Techniques: Infection, Quantitative RT-PCR, Expressing, Comparison, Transfection
Journal: bioRxiv
Article Title: SARS-CoV-2 Nsp15 antagonizes the cGAS-STING-mediated antiviral innate immune responses
doi: 10.1101/2024.09.05.611469
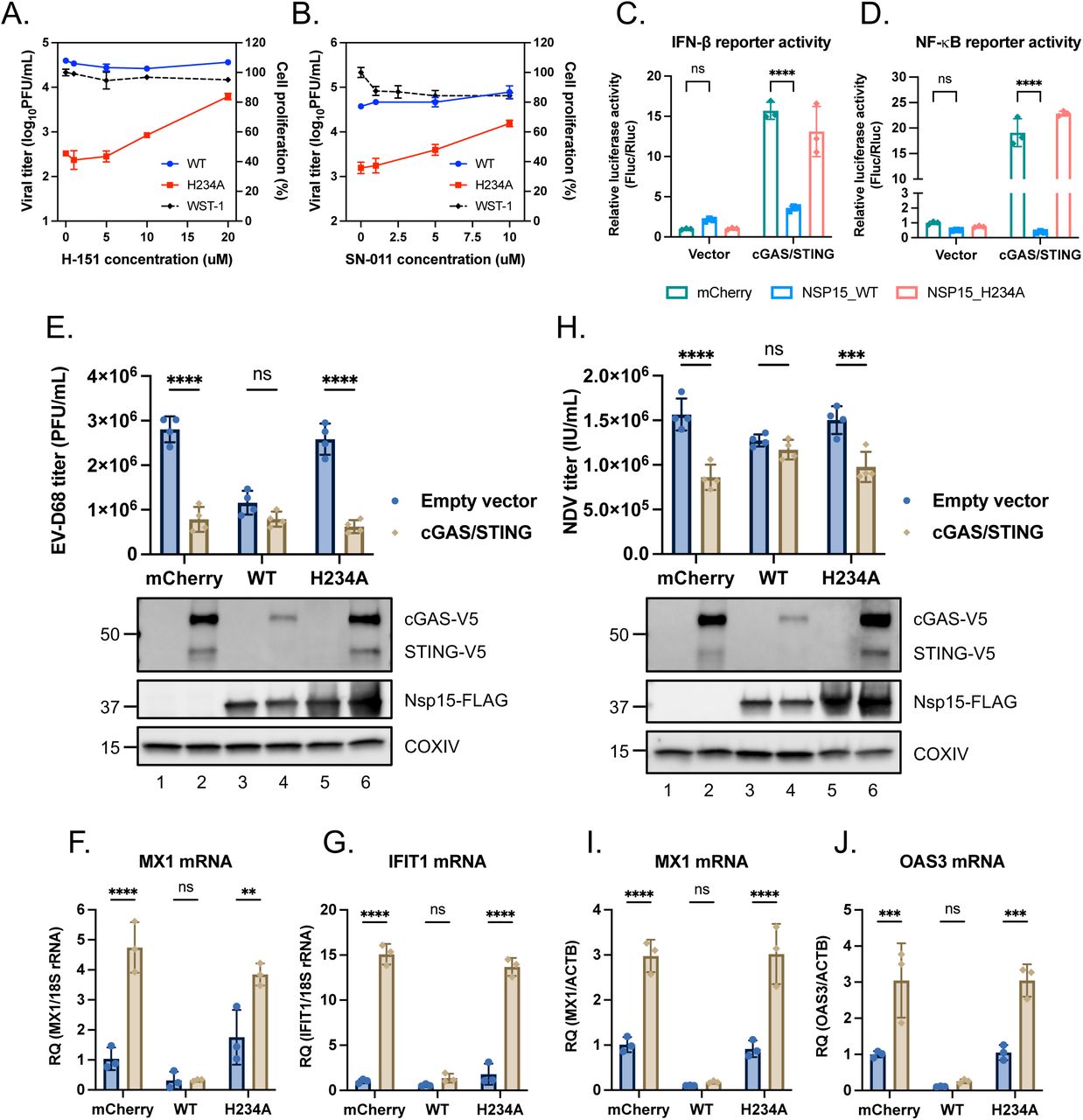
Figure Lengend Snippet: (A and B) A549-ACE2 cells were pretreated with H-151 or SN-011 for 16 h prior to viral infection. Cells were then infected with Nsp15 WT (blue line) and Nsp15 H234A (red line) rSARS-CoV-2 at MOI of 1 under H-151 or SN-011 treatment for 48 h. Supernatants harvested for viral titration by plaque assay. Cell viability determined by WST-1 assay after incubating with H-151 or SN-011 for 72 h (black dot line). Data are mean ± SD (n = 3). (C and D) HEK293T cells were co-transfected with IFN-β promoter reporter plasmid (C) or NF-κB responsive element reporter plasmid (D), Renilla control plasmid plus the indicated plasmids containing empty vector, cGAS/STING, mCherry and WT and H234A Nsp15 for 48 h. Relative luciferase activity was performed by use of Dual-Glo Luciferase System. Data are mean ± SD (n = 3) and analyzed by two-way ANOVA with Dunnett’s multiple comparison test. (E, F, G) HEK293T transfected with different combinations of plasmids were infected with EV-D68 at MOI of 0.1 for 48 h. Culture supernatants applied for viral titer quantification by plaque assay (E, upper bar graph). Cell lysates collected for protein level determination (E, lower blot graph) and total RNA collected for RT-qPCR (F, G). (H, I, J) HEK293T transfected with different combinations of plasmids were infected with NDV-EGFP at MOI of 1 for 48 h. Culture supernatants applied for viral titer quantification by focus forming assay targeting EGFP (H, upper bar graph). Cell lysates collected for protein level determination (H, lower blot graph) and total RNA collected for RT-qPCR (I, J). Relative target RNA level normalized with that of 18S rRNA or ACTB was shown. Data are mean ± SD (n = 3 or 4). and analyzed by two-way ANOVA with Šídák multiple comparison test. ** P ≤0.01; *** P ≤0.001; **** P ≤0.0001; ns, not significant.
Article Snippet: BHK-21-hACE2 cell, a derivative of BHK-21 cell (ATCC, CCL-10), was established by transduction of lentiviral particle bearing
Techniques: Infection, Titration, Plaque Assay, WST-1 Assay, Transfection, Plasmid Preparation, Control, Luciferase, Activity Assay, Comparison, Quantitative RT-PCR, Focus Forming Assay

Journal: medRxiv
Article Title: The Contrasting Role of Nasopharyngeal Angiotensin Converting Enzyme 2 ( ACE2 ) Expression in SARS-CoV-2 Infection: A Cross-Sectional Study of People Tested for COVID-19 in British Columbia
doi: 10.1101/2020.11.23.20237206
Figure Lengend Snippet: The other does not include the transmembrane domain (pink) and lays nested between two motifs known to interact with the SARS-CoV receptor binding domain (blue). The measure of soluble ACE2 represents normalized expression of the pink to yellow target, or the proportion of non-transmembrane ACE2 to transmembrane ACE2 .
Article Snippet: The
Techniques: Binding Assay, Expressing

Journal: medRxiv
Article Title: The Contrasting Role of Nasopharyngeal Angiotensin Converting Enzyme 2 ( ACE2 ) Expression in SARS-CoV-2 Infection: A Cross-Sectional Study of People Tested for COVID-19 in British Columbia
doi: 10.1101/2020.11.23.20237206
Figure Lengend Snippet: Boxplots of transmembrane ACE2 expression by nine-year age categories in COVID-19-negative participants between the ages of 19 and 98 (n=198); boxes represent the Q1-Q3 interquartile range, whiskers represent 1.5x the Q1 or Q3 and horizontal lines the median transmembrane ACE2 expression by age category. Nine-year categories were selected to optimize the distribution of observations between groups. Participants who tested negative younger than 19 or older than 98 were excluded based on n<10 observations per age group. No difference was detected in mean transmembrane ACE2 expression among age categories (ANOVA, P=0·092).
Article Snippet: The
Techniques: Expressing

Journal: medRxiv
Article Title: The Contrasting Role of Nasopharyngeal Angiotensin Converting Enzyme 2 ( ACE2 ) Expression in SARS-CoV-2 Infection: A Cross-Sectional Study of People Tested for COVID-19 in British Columbia
doi: 10.1101/2020.11.23.20237206
Figure Lengend Snippet: Gene expression is portrayed in kernel density plots stratified by COVID-19 test result. Probability densities of relative host gene expression are shown by positive (red, transmembrane ACE2 ; blue, soluble ACE2 ; yellow, TMPRSS2 ) and negative COVID-19 test results (grey). Levene’s test was used to detect non-equal variance in gene expression for all host targets between COVID-19-negative and -positive participants. A two-tailed, t-test was used to examine mean difference in host gene expression by COVID-19 test result assuming unequal variance: transmembrane ACE2 (P=1·2e-4), soluble ACE2 (P<0·0001) and TMPRSS2 (P<0·0001).
Article Snippet: The
Techniques: Gene Expression, Two Tailed Test

Journal: medRxiv
Article Title: The Contrasting Role of Nasopharyngeal Angiotensin Converting Enzyme 2 ( ACE2 ) Expression in SARS-CoV-2 Infection: A Cross-Sectional Study of People Tested for COVID-19 in British Columbia
doi: 10.1101/2020.11.23.20237206
Figure Lengend Snippet: The level of transmembrane ACE2 (red), soluble ACE2 (blue) and TMPRSS2 (black) expression in positive cases of COVID-19 and its relationship with SARS-CoV-2 RNA is shown in Log 10 genome equivalents per millilitre (GE/mL). Host gene expression was measured by quantitative real-time PCR and normalized to expression of the housekeeping gene GAPDH by the (2 -ΔΔCt ) method. Soluble ACE2 was defined as absolute relative expression between the Hs01085333_m1 (ThermoFisher) and HS01085340_m1 (ThermoFisher) gene targets . Simple linear regression was used estimate correlations, the reported P values are for the B eta-coefficients of host gene expression.
Article Snippet: The
Techniques: Expressing, Gene Expression, Real-time Polymerase Chain Reaction

Journal: medRxiv
Article Title: The Contrasting Role of Nasopharyngeal Angiotensin Converting Enzyme 2 ( ACE2 ) Expression in SARS-CoV-2 Infection: A Cross-Sectional Study of People Tested for COVID-19 in British Columbia
doi: 10.1101/2020.11.23.20237206
Figure Lengend Snippet: Soluble ACE2 expression was categorized into low, mean and high levels to demonstrate the relationship. The low category (grey) represents soluble ACE2 expression two standard deviations below the mean expression. The mean category (orange) codes for the mean soluble ACE2 expression, zero standard deviations. The high category (blue) indicates soluble ACE2 expression two standard deviations above mean expression. Shaded areas represent 95% confidence intervals, solid lines represent B coefficients from multiple linear regression.
Article Snippet: The
Techniques: Expressing